Free-living amoebae isolated in the Central African Republic: epidemiological and molecular aspects
Alain Farra, Claudine Bekondi, Vianney Tricou, Jean Robert Mbecko, Antoine Talarmin
Corresponding author: Alain Farra, Institut Pasteur de Bangui, Bangui, Republique Centrafricaine 
Received: 03 Feb 2016 - Accepted: 19 Dec 2016 - Published: 01 Feb 2017
Domain: Public Health
Keywords: Free-living amoebae, tetramitus, CAR
©Alain Farra et al. Pan African Medical Journal (ISSN: 1937-8688). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Alain Farra et al. Free-living amoebae isolated in the Central African Republic: epidemiological and molecular aspects. Pan African Medical Journal. 2017;26:57. [doi: 10.11604/pamj.2017.26.57.9021]
Available online at: https://www.panafrican-med-journal.com//content/article/26/57/full
Original article 
Free-living amoebae isolated in the Central African Republic: epidemiological and molecular aspects
Free-living amoebae isolated in the Central African Republic: epidemiological and molecular aspects
Alain Farra1,&, Claudine Bekondi1, Vianney Tricou1, Jean Robert Mbecko1, Antoine Talarmin2
1Institut Pasteur de Bangui, Bangui, Republique Centrafricaine, 2Institut Pasteur de Guadeloupe, France
&Corresponding author
Alain Farra, Institut Pasteur de Bangui, Bangui, Republique Centrafricaine
Among the many species of free-living amoebae infecting humans, only Naegleria fowleri, a few species of Acanthamoeba, Balamuthia mandrillaris recently Sappinia diploďdea and Paravahlkampfia Francina are responsible for human diseases especially deadly encephalitis outside of Acanthamoeba keratitis related . In the Central African Republic (CAR), no studies have previously been conducted about free amoebae and no suspicious cases of encephalitis or amoebic keratitis was reported even though the ecosystem supported the proliferation of these microorganisms. The objective of this study was to identify free-living amoebae present in CAR and to define the molecular characteristic. Bathing sites and cerebrospinal fluid from patients died of bacterial meningitis untagged were explored by culture and PCR and the amplicons were sequenced which allowed to characterize the species found. Only species of the genus Tetramitus, namely T. Entericus, T. waccamawensis and T.sp similar to those already described in the world and not pathogenic for humans were found in bathing sites, the cerebrospinal fluid meanwhile remained negative. Although no pathogen species such as Naegleria fowleri or species of Acanthamoeba have been isolated, this study worth pursuing because this investigation was very limited in space because of the insecurity in the country.
Free-living amoebae, unlike parasitic amoebae, complete their entire cycle in nature and do not require a host [1]. Some free-living amphizoic amoebae can, however, accidentally infect humans and cause neurological, ocular and cutaneous infections [2, 3]. The main organisms involved are Naegleria, Acanthamoeba, Balamuthia, several amoebae of the genus Sappinia (S. diploidea, S. pedata) and a species of the genus Paravahlkampfia, P. francina, which was recently incriminated in cases of encephalitis [4-6]. Some of the cases of encephalitis were opportunistic infections in immunodepressed individuals and consisted of granulomatous encephalitis due to Acanthamoeba and Balamuthia, which evolves chronically and is usually fatal. In contrast, primary amoebic meningoencephalitis due to Naegleria fowleri is an acute condition in healthy children and adults, manifesting several days after infection and rapidly evolving to severe disease in the absence of early treatment. In the Central African Republic (CAR), the presence of free-living amoebae has not been studied, and no suspected cases of primary amoebic meningoencephalitis have been reported, although cases have been reported in Nigeria, Zambia and South Africa [7, 8]. The presence in the CAR of hot springs and numerous warm-water lakes with abundant organic matter would indicate that such organisms might exist there also. The objective of this study was to identify and characterize the free-living amoebae that are present and to determine whether they include N. Fowleri and try to assess the risk of the population.
Context geographic: the CAR is a landlocked country in the heart of the African continent in an intertropical zone. It covers an area of 623 000 km2 and is bordered on the east by the Republic of Sudan and South Sudan, on the west by Cameroon, on the south by the Democratic Republic of the Congo and the Congo, and on the north by Chad. Its geographical position results in a hot continental climate, with two seasons: a dry season between November and April and a rainy season between May and October. The country has numerous lakes, ponds, rivers and hot springs, in which people swim. We obtained our samples from some of these water bodies, but the insecurity associated with the current military-political situation limited our study to accessible areas near Bangui.
Sample Collection: we took samples from 32 sites between 4 April and 23 May 2012 in Bangui and in three directions within a radius of 150 km around the city: Bangui-Damara-Mbourouba, Bangui-Boali and Bangui-Mbaďki-Mbata (Figure 1). The sites contain rivers, lakes, ponds, pools and a few functioning hotel swimming pools in Bangui. The hot springs at Dessikou in Dékoua in the centre of the country and at Nzako near Bria in the east could not be sampled because of security problems. At each site, we noted the temperature and pH of the water and the GPS coordinates. Samples were taken in duplicate for incubation at 37°C and 44 ° C and consisted of 500 ml of water, algae and swab samples from swimming pools. Sediments could be collected only from swimming pools because the bottom of all the other water bodies was mud. The samples were sent to the Institut Pasteur and cultured within 6 h of collection. With the approval of the Ethics Committee, we examined 20 samples of purulent cerebrospinal fluid (CSF) from the biological specimen library at the Institut Pasteur by culture and by PCR to identify any amoebae. These samples were from HIV-negative patients who had died of acute meningoencephalitis for which no bacterial cause had been found by culture or by molecular biology.
Isolation of Amoebae: the water samples were filtered with a Millipore® model DOA U152-BN on cellulose nitrate filters with a pore width of 1.2 µm. The filters were cut into eight pieces in a class II microbiological safety cabinet and returned to a non nutrient agar previously flooded with a suspension of Escherichia coli, which is used as a nutrient by amoebae. The algae and swab samples were placed directly on the culture medium under the same conditions. The CSF samples were seeded in three spots on the culture medium. All the dishes were then sealed with Parafilm® and incubated at 37°C, at which temperature all free-living amoebae grow, an at 44°C to select for certain species of N. fowleri. Reading cultures was made first to the eye and the inverted phase contrast microscope and this every day for at least 5 days. To the naked eye, the colonies of amoebae in the form of an opaque white halo around the deposit to visible white light. They were surrounded with felt on the bottom of the box and then observed under a microscope with a X20 objective which allows to view trophozoďtes and sometimes cysts around the deposits. When a culture was positive, a section of agar containing amoebae was cut out with an öse in the safety cabinet, returned to fresh culture medium and incubated as above.
DNA Extraction, PCR Amplification and Sequencing: DNA was extracted from the cultures with a DNA Mini-Kit® (50) (Qiagen) and quantified before PCR with a Qubit 2.0 fluorimeter according to the manufacturer´s instructions. A concentration of 10-20 ng/µl was used for PCR. As morphological identification is difficult, species were identified exclusively by molecular biology with PCR, followed by amplicon sequencing. Three pairs of primers were used respectively for summer generic looking amoebae (JITSFW: 5'-GTCTTCGT AGGTGAACCTGC-3', JITSRV : 5'-CCGCTTACTGATATGCTTAA-3') Naegleria sp. (ITSFW : 5'-AACCTGCGTAGGGATCATTT-3', ITSRW: 5'-TTTCCTCCCCTTATTAATAT-3' ) and N. fowleri. (NFITSFW: 5'-TGAAAACCTTTTTTCCATTTACA-3', NFITSRV: 5'-AATAAAAGAT TGACCATTTGAAA-3'). The sequencing of the amplicons allowed the diagnosis of species. The amplicons were purified with a QIAquick PCR purification kit (Qiagen) and then sent to GATC Biotech in Germany for sequencing. Analysis with the Blast 2.0 program identified separate species. Sequences were aligned with the Clustal W2 program, and the phylogenetic tree was constructed with MEGA 5.2.2.
The temperature of the water at the sampled sites did not exceed 40°C, and that at 47% of the sites was ≥ 30°C (Table 1,Table 2, Table 3). Samples from only eight of the 32 sites were positive on culture. Five of the cultures were from algae and three from filtered water, but the samples of algae and water were from different sites. Only the cultures incubated at 37°C for 72 h were positive; all those incubated at 44°C were sterile, as were those of CSF. DNA of free-living amoebae was identified by PCR in seven of the eight positive cultures with JITS primers. Naegleria DNA was found in two samples (ITS primers), but N. fowleri was not identified (Table 4). Furthermore, no amoebic DNA was found in CSF, and sequencing showed no Naegleria species, only the species of Tetramitus namely T. waccamawensis, T. entericius and .T. SP. were identified with the analysis of the sequences (Table 4). The physicochemical characteristics of the various sites did not offer any clues, as the species found were all of the same genus (Table 1,Table 2, Table 3). The phylogenetic study showed that the species found in the CAR were identical to those found in Australia and the USA and were very similar to other Tetramitus isolated elsewhere in the world (Figure 2).
This preliminary search for free-living amoebae in the aquatic environment is the first of its kind in the CAR. It was limited spatially because of the lack of security in the country. The temperature of the water in most of the bodies studied was 25-35°C, and none had a temperature superior at 40°C. No Naegleria or Acantamoeba species was isolated, even though these species are ubiquitous and they are probably present in the CAR. The results show that the conditions at certain sites are favourable for the growth of N. fowleri, with abundant organic matter and a temperature superior at 30°C; the water at two sites was even superior at 35°C [5, 8, 9]. A case of infection with N. fowleri described in Guadeloupe occurred after bathing in water at 27°C, and N. fowleri has been found in water at 26.9-34.9°C [10, 11]. Almost all the sites studied were compatible with the presence of this amoeba, which would justify continuation of this study. It will therefore be extended to other sites and particularly hot springs, once the security situation improves. In the cases of primary amoebic meningoencephalitis seen in Madagascar and Guadeloupe, the amoebae were visible in fresh CSF, and PCR of frozen CSF showed the presence of N. fowleri [10, 12]. Although our samples were kept at -20°C under good conditions, PCR showed no amoebae. A systematic prospective study of purulent CSF samples with no bacterial cause should be undertaken, with careful direct examination and generic PCR to detect amoebae. The only amoeba species that were isolated belonged to the Tetramitus genus in the Vahlkampfidae family. Currently, 17 species have been associated with disease in humans [13]. T. waccamawensis was previously classified as Learamoeba waccamawensis according to the criteria of Sawyer et al. [14], but a molecular study by Brown and De Jonckheere led to its reclassification in the genus Tetramitus on the basis of 98.7% homology with the amoebae of this genus [15]. Except for two species, T. jugosus in the marine environment and T. thermacidophilus which develops at a pH of 1.2-5 and at temperatures up to 54°C, Tetramitus are usually isolated from freshwater with abundant organic matter, as in the CAR where this amoeba was isolated [16-18]. The two strains identified by sequencing, T. entericius and T. waccamawensis are similar to the two species already described [13]. The unidentified strain is also similar to these two species; a further study will be conducted to identify this strain.
As expected, free-living amoebae are present in freshwater in the CAR. They all belong to the Tetramitus genus and are not pathogenic to humans. No species of the genera Naegleria or Acanthamoeba was isolated, either from the environment or from purulent CSF samples with no bacterial agent. This study should be extended to other sites and particularly to the Dessikou and Nzako hot springs, which could favour the proliferation of Naegleria . Better understanding of the ecology and epidemiology of free-living amoebae in the CAR, both in the environment and in HIV-infected patients, is needed, in order to raise awareness about these neglected infections, which are usually fatal, and to put in place prophylactic measures.
What is known about this topic
- Free-living amoebae and roles in human pathologies;
- Meningoencephalitis caused by Naegleria fowleri;
- Amoebic keratitis caused by acanthamoeba.
What this study adds
- Free-living amoebae present in RCA;
- Ecosystem favourable to the presence of Naegleria fowleri;
- Possible diagnosis at the Institute Pasteur of Bangui.
The authors declare no competing interests.
Alain Farra was the principal investigator, working from levies on sites, cultivation, the realization of the PCR, the sequence analysis as well as the drafting of this article. Alain Farra was accompanied by Mr Jean Robert Mbecko for sampling and it has also actively participated in the realization of the manipulations in the laboratory. Claudine Bekondi provided the CSF of its service for the exploration of the amoebae. Vianney Tricou performed the phylogenetic analysis, Antoine Talarmin has designed the research project and made it operational as well in Guadeloupe, Madagascar and Central African Republic through a concerted inter Pasteur.
We thank Mirdad Kazanji former Director of the Institut Pasteur in Bangui for the translation of the English text, the Drs Mirna Moussa Scientist in the Pasteur Institute of Guadeloupe and Johan De Jonckheere from the Brussels Institute of Public Health for their contribution to the realization of this study. This work has benefited particulary from the expertise of Johan De Jonckeere to whom we send photos of the electrophoresis of PCR products and sequences of amoebae for confirmation.
Table 1: results of culture and PCR of samples in Bangui
Table 2:
results of culture and PCR of samples on axis 1 and 2
Table 3: results of culture and PCR of samples on axis 3
Table 4: results PCR and species identified by sites
Figure 1: map of the CAR,
showing the directions chosen for sampling and the security situation in the
country; This map was made available to all partners in health by the WHO are
indicated the different axes (A1: Axis Bangui-Damara-Mbourouba; A2: Axis Bangui-Boali;
A3: Axis Bangui-Mbaďki-Mbata) and Dessikou and Nzako cities that could not be
investigated
Figure 2: phylogeny of amoebae found in CAR
- Aubry P. Infections ŕ amibes libres. Actualités 2010. Texte mis ŕ jour le 26/10/2011. Google Scholar
- Page FC. A New Key to Freshwater and Soil Gymnamoebae. Freshwater Biological Association, Ambleside, Cumbria, England. (1988); 122pp. PubMed | Google Scholar
- Visvesvara GS, Moura H, Schuster FL. Pathogenic and opportunistic free-living amoebae: Acanthamoeba spp, Balamuthia mandrillaris, Naegleria fowleri, and Sappinia diploidea. FEMS Immunol Med Microbiol. 2007 Jun;50(1):1-26. PubMed | Google Scholar
- Heggie TW. Swimming with death: Naegleria fowleri infections in recreational waters. Travel Med Infect Dis. 2010 Jul;8(4):201-6. PubMed | Google Scholar
- Lorenzo-Morales J, Martín-Navarro M C, Martínez-Carretero E, Jose E Pinero, Basilio Valladares B. Encephalitis Due to Free Living Amoebae: An Emerging Issue in Human Health. In: Tkachev S, editor. Non-Flavivirus Encephalitis: InTech; 2011. Google Scholar
- Visvesvara GS, Sriram R, Qvarnstrom Y, Bandyopadhyay K, Da Silva AJ, Pieniazek NJ et al. Paravahlkampfia francinae n sp masquerading as an agent of primary amoebic meningoencephalitis. J Eukaryot Microbiol. 2009 Jul-Aug;56(4):357-66. PubMed | Google Scholar
- Lawande RV, Macfarlane JT, Weir WR, Awunor-Renner C. A case of primary amebic meningoencephalitis in a Nigerian farmer. Am J Trop Med Hyg. 1980 Jan;29(1):21-5. PubMed | Google Scholar
- Schoeman CJ, van der Vyver AE, Visvesvara GS. Primary amoebic meningo-encephalitis in southern Africa. J Infect. 1993 Mar;26(2):211-4. PubMed | Google Scholar
- Tyndall RL, Ironside KS, Metler PL, Tan EL, Hazen TC, Fliermans CB. Effect of thermal additions on the density and distribution of thermophilic amoebae and pathogenic Naegleria fowleri in a newly created cooling lake. Appl Environ Microbiol. 1989 Mar;55(3):722-32. PubMed | Google Scholar
- Nicolas M, De Jonckheere JF, Pernin P, Bataille H, Le Bris V, Herrmann-Storck C. Molecular diagnosis of a fatal primary amoebic meningoencephalitis in Guadeloupe (French West Indies). Bull Soc Pathol Exot. 2010 Feb;103(1):14-8. PubMed | Google Scholar
- Moussa M, De Jonckheere JF, Guerlotté J, Richard V, Bastaraud A, Romana M et al. Survey of Naegleria fowleri in geothermal recreational waters of Guadeloupe (French West Indies). PLoS One. 2013;8(1):e54414. PubMed | Google Scholar
- Jaffar-Bandjee MC, Alessandri JL, Molet B, Clouzeau J, Jacquemot L, Sampériz S et al. Primary amebic meningoencephalitis: 1st case observed in Madagascar. Bull Soc Pathol Exot. 2005 Apr;98(1):11-3. PubMed | Google Scholar
- De Jonckheere JF, Brown S. Isolation of a vahlkampfiid amoeba from a contact lens: Tetramitus ovis (Schmidt, 1913) n comb. European Journal of Protistology. 2005 4/29/;41(2):93-7. PubMed | Google Scholar
- Sawyer TK, Nerad TA, Cahoon LB, Nearhoof JE. Learamoeba waccamawensis, N G, N Sp (Heterolobosea: Vahlkampfiidae), a new temperature-tolerant cyst-forming soil Amoeba. Journal of Eukaryotic Microbiology. 1998;45(3):260-4. PubMed | Google Scholar
- Brown S, De Jonckheere JF. A reevaluation of the amoeba genus Vahlkampfia based on SSUrDNA sequences. European Journal of Protistology. 1999/02/25;35(1):49-54. PubMed | Google Scholar
- Read LK, Margulis L, Stolz J, Obar R, Sawyer TK. A new strain of paratetramitus jugosus from laguna figueroa, Baja California, Mexico. Biological bulletin. 1983;165(1):241-64. PubMed | Google Scholar
- Enzien M, McKhann HI, Margulis L. Ecology and life history of an amoebomastigote, Paratetramitus jugosus, from a microbial mat: new evidence for multiple fission. Biological Bulletin. 1989 Aug;177(1):110-29. PubMed | Google Scholar
- Baumgartner M, Eberhardt S, De Jonckheere JF, Stetter KO. Tetramitus thermacidophilus n sp, an amoeboflagellate from acidic hot springs. J Eukaryot Microbiol. 2009 Mar-Apr;56(2):201-6. PubMed | Google Scholar
Search
This article authors
On Pubmed
On Google Scholar
Citation [Download]
Navigate this article
Similar articles in
Key words
Tables and figures
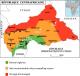 Figure 1: map of the CAR, showing the directions chosen for sampling and the security situation in the country; This map was made available to all partners in health by the WHO are indicated the different axes (A1: Axis Bangui-Damara-Mbourouba; A2: Axis Bangui-Boali; A3: Axis Bangui-Mbaïki-Mbata) and Dessikou and Nzako cities that could not be investigated
Figure 1: map of the CAR, showing the directions chosen for sampling and the security situation in the country; This map was made available to all partners in health by the WHO are indicated the different axes (A1: Axis Bangui-Damara-Mbourouba; A2: Axis Bangui-Boali; A3: Axis Bangui-Mbaïki-Mbata) and Dessikou and Nzako cities that could not be investigated












